Links to this work:
Summary
This thesis presents a UML extension with its concrete syntax and written template to document bioinformatics workflows. The produced artefacts were evaluated and validated in cooperation with three facilities and seven bioinformaticians following a data science methodology. The results of this work are our contribution to the bioinformatics domain and the ongoing research: Optimized Bioinformatics Workflows from Requirements Engineering of Solution Specifications.
Methodology
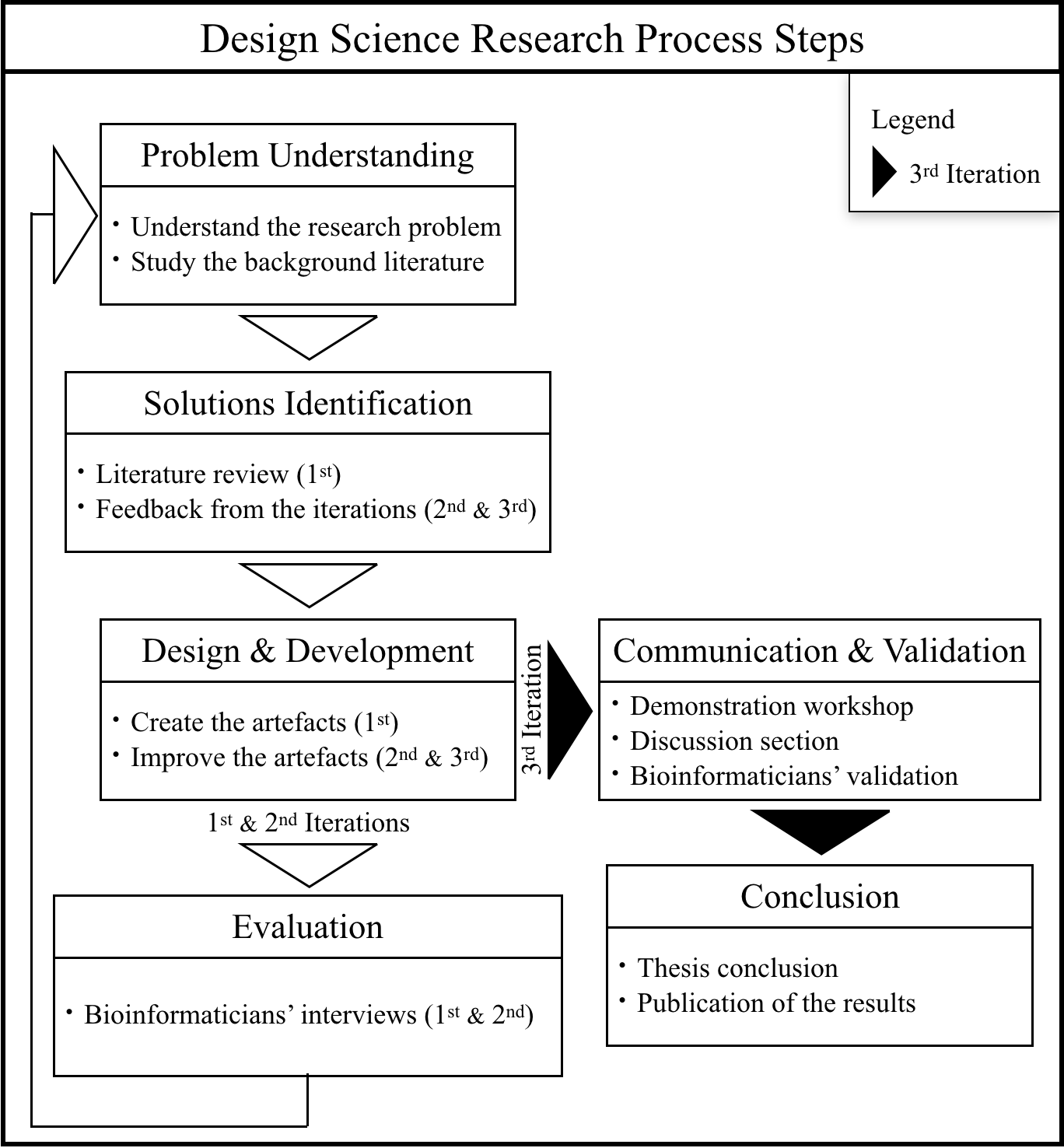
First Iteration
- Questionnaire
- Metamodel
- WDST
- Concrete Syntax 1
- Concrete Syntax 2
- Concrete Syntax example 1
- Concrete Syntax example 2
- Legend for the examples
Interaction Materials:
Second Iteration:
- Enhancements done in the Meta-model
- WDST changes
Improvements:
- Questionnaire
- Metamodel
- Concrete Syntax
- Concrete Syntax
- WDST
- Concrete Syntax Example
- Legend for the Example
Interaction Materials:
Third Iteration:
- Enhancements done in the Concrete Syntax
- WDST changes
Improvements:
- Questionnaire
- Final Metamodel
- Concrete Syntax
- Final WDST
- Concrete Syntax Example
- WDST Example
- Workshop presentation
Study Materials:
- Interview Codebook
- Concrete syntax evaluation findigns
- WDST evaluation findigns
- Mentimeter Validation